What are protein interaction networks?
Protein interaction networks represent all of the interactions between a particular protein or multiple proteins [1]. The study of protein-protein interactions allows for better understanding of protein function at both an individual and a global level. When studying proteins of unknown functions, protein interaction networks can provide key insight into that protein's function, location, significance, and relation to disease progression. One way to investigate protein interaction networks is through the database called STRING, which compiles data from multiple research databases while also making predictions based on text-mining in order to construct a protein interaction network for your desired protein [2].
Protein interaction networks represent all of the interactions between a particular protein or multiple proteins [1]. The study of protein-protein interactions allows for better understanding of protein function at both an individual and a global level. When studying proteins of unknown functions, protein interaction networks can provide key insight into that protein's function, location, significance, and relation to disease progression. One way to investigate protein interaction networks is through the database called STRING, which compiles data from multiple research databases while also making predictions based on text-mining in order to construct a protein interaction network for your desired protein [2].
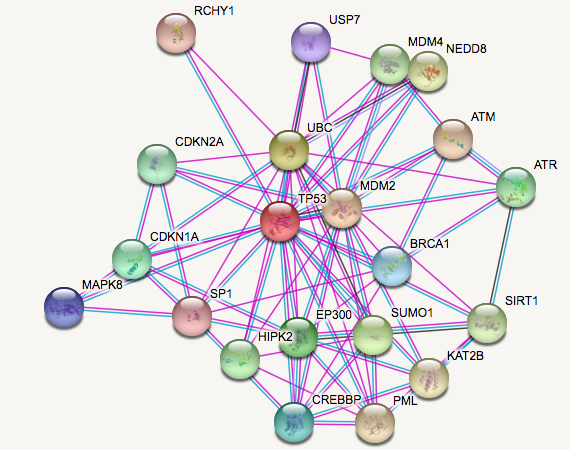
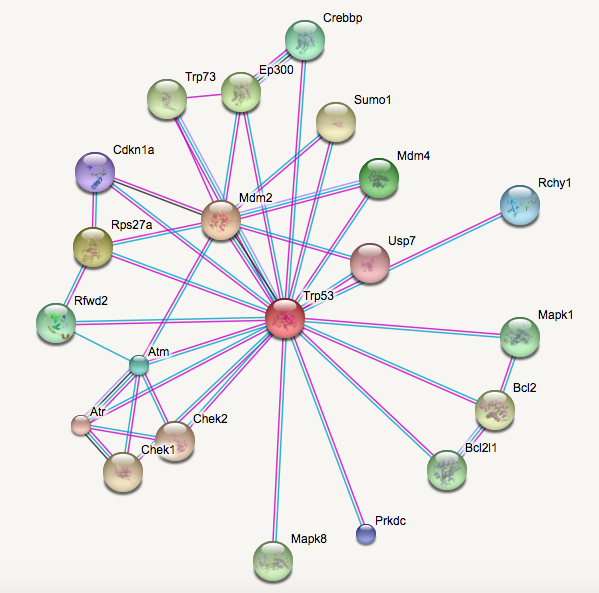
What do the interaction networks for TP53 look like in humans and mice?
Conclusion:
Even though human TP53 protein and mouse Trp53 are pretty similar in regards to domains and post-translational modifications, their protein interaction networks are notably different. One reason for this difference could be because the protein homologs for other proteins in the mouse model may be significantly more different than in humans compared to TP53. However, these differences could also be used to explore protein-protein interactions related specifically to ovarian cancer since the disease can spontaneous occur in humans but does not spontaneously occur in mice.
Even though human TP53 protein and mouse Trp53 are pretty similar in regards to domains and post-translational modifications, their protein interaction networks are notably different. One reason for this difference could be because the protein homologs for other proteins in the mouse model may be significantly more different than in humans compared to TP53. However, these differences could also be used to explore protein-protein interactions related specifically to ovarian cancer since the disease can spontaneous occur in humans but does not spontaneously occur in mice.
Resources:
[1] Vazquez, A. (2010). Protein Interaction Networks. CRC Press/Taylor & Francis. Retrieved April 11, 2017. https://www.ncbi.nlm.nih.gov/books/NBK56024/#ch8_sec1
[2] String Consortium. (2017). STRING. Retrieved April 11, 2017. http://string-db.org/cgi/input.pl UserId=lLpPTAafPWFW&sessionId=mfizZh27dqm9&input_page_show_search=on
[1] Vazquez, A. (2010). Protein Interaction Networks. CRC Press/Taylor & Francis. Retrieved April 11, 2017. https://www.ncbi.nlm.nih.gov/books/NBK56024/#ch8_sec1
[2] String Consortium. (2017). STRING. Retrieved April 11, 2017. http://string-db.org/cgi/input.pl UserId=lLpPTAafPWFW&sessionId=mfizZh27dqm9&input_page_show_search=on